This page is intended as a resource. Click here to read about our engineering services, including converting any antibody into a fragment.
There are a range of applications in which Fc mediated effects are not required and are even undesirable. A common solution for applications where the antibody is only being used to block a signalling molecule or receptor is the use of antibody fragments that lack the Fc domain (1). This also helps to reduce the other main failure of therapeutic antibodies, namely the lack of delivery, which is especially true for anti-cancer antibodies. Solid tumours have substantial physical barriers often preventing the penetration of antibodies to the centre and resulting in reduced therapeutic effects (2). The use of smaller fragments enables deeper penetration with the affinity of the antibody also being critical and if it is too high this will restrict its ability to penetrate a tumour (3). Fragments are also useful in imaging and cancer therapy, where a long serum half-life mediated by Fc interaction with the FcRn receptor results in poor contrast and in the case of radiolabelled antibodies fast clearance from the circulation via the kidney is also advantageous to reduce prolonged exposure (4). Antibody fragments that have been engineered to be multimeric are of use when targeting multiple disease associated antigens. This approach and others will be described in more detail in the bispecifics section.
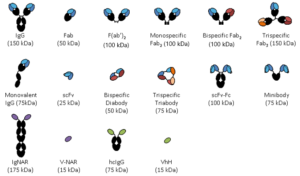
Over the past three decades antibodies have been dissected into smaller antigen binding fragments, initially by proteolysis and later by genetic engineering to produce mono or multivalent fragments. The image above shows a selection of the antibody fragments that have been engineered. These are mainly based on either Fab fragments or the single chain Fv (scFv) as building blocks. The Fv portion of an IgG, consisting of the VH and VL domains, is the smallest fragment that maintains the full binding capacity of the intact antibody. However, Fv domains tend to have limited stability due to the dissociation of the domains and thus a peptide linker was introduced between the VH and VL domains to create the scFv (5, 6). ScFvs naturally multimerize (7) and this can be controlled by linker length creating dimeric, trimer or tetrameric forms (8, 9). For this reason scFvs have become one of the main building blocks of antibody fragments.
When mouse single VH domains were shown to be functional (10) it was proposed that because of their small size they could potentially target cryptic epitopes that have evolved in many pathogenic viruses to contain narrow cavities that bind to target receptors but are poorly accessible to intact antibodies. However, they rarely retain the affinity of their parent antibody and are poorly soluble and prone to aggregation (10). Discovery of two types of organisms, the camelids and cartilaginous fish, that have evolved high affinity single variable domains (called the VhH domain in camelids (11) and the V-NAR domain in sharks (12)) mounted on an Fc equivalent constant domain framework resparked interest in this area. Both of these domains contain long CDR loops, often larger than those observed in conventional murine and human antibodies. Thus these are able to penetrate cavities in target antigens such as active sites (13) and canyons in viral and infectious disease biomarkers (14, 15) that may be inaccessible to the more planar binding associated with human or murine antibodies. At least two biotech companies have been founded on single domain antibody based technologies, Domantis (now part of GlaxoSmithKline) and Ablynx, with these molecules now starting to show promise in clinical trials (16).
To the best of our knowledge the smallest reported antibody fragment is a 3 kDa construct consisting of one CDR from the heavy chain and one from the light chain separated by a framework region (17). It was proposed that separation of two CDRs by a framework region would allow loops to assume conformation similar to that of the parental antibody. This was indeed shown to be true, although the constructs bound their antigen with only 1-10% of the affinity of the parent antibody as may have been expected given the reduced number of CDRs.
Despite the many advantages discussed above one of the main drawbacks of most antibody fragments for use as therapeutics is the lack of an Fc domain. This prevents FcRn-mediated recycling and thus infers a greatly reduced half-life, typically in the region of hours to days as opposed to weeks for a typical IgG. This has led to more recent efforts to either retain this natural property of IgG or reinstate it by further engineering. Examples of the former approach include the one-armed MetMAb developed by Genentech, which contains a heterodimeric Fc with only one Fab arm (18), or a monovalent IgG (also known as a half-antibody) developed in a number of different ways independently by Genmab (19), MedImmune (20), Pfizer (21) and the NIH (22). An example of the latter approach includes the use of engineered albumin binding domains fused to the antibody fragment to infer an albumin-like half-life (23)
A complete review of all antibody fragments in development is beyond the scope of this resource. For further information on antibody fragments we recommend reviews from Holliger and Hudson (1), Cuesta et al (24) and Nelson (25).
<< Humanization Fc engineering >>
References
- Holliger, P., and Hudson, P.J. (2005). Engineered antibody fragments and the rise of single domains. Nat. Biotechnol. 23, 1126–1136.
- Christiansen, J., and Rajasekaran, A.K. (2004). Biological impediments to monoclonal antibody-based cancer immunotherapy. Mol. Cancer Ther. 3, 1493–1501.
- Adams, G.P., Schier, R., McCall, A.M., Simmons, H.H., Horak, E.M., Alpaugh, R.K., Marks, J.D., and Weiner, L.M. (2001). High affinity restricts the localization and tumor penetration of single-chain fv antibody molecules. Cancer Res. 61, 4750–4755.
- Brekke, O.H., and Løset, G.Å. (2003). New technologies in therapeutic antibody development. Curr Opin Pharmacol 3, 544–550.
- Bird, R.E., Hardman, K.D., Jacobson, J.W., Johnson, S., Kaufman, B.M., Lee, S.M., Lee, T., Pope, S.H., Riordan, G.S., and Whitlow, M. (1988). Single-chain antigen-binding proteins. Science 242, 423–426.
- Huston, J.S., Levinson, D., Mudgett-Hunter, M., Tai, M.S., Novotný, J., Margolies, M.N., Ridge, R.J., Bruccoleri, R.E., Haber, E., and Crea, R. (1988). Protein engineering of antibody binding sites: recovery of specific activity in an anti-digoxin single-chain Fv analogue produced in Escherichia coli. Proc. Natl. Acad. Sci. U.S.A. 85, 5879–5883.
- Wilkinson, I.C., Hall, C.J., Veverka, V., Shi, J.Y., Muskett, F.W., Stephens, P.E., Taylor, R.J., Henry, A.J., and Carr, M.D. (2009). A high resolution NMR-based model for the structure of a ScFv-IL-1b Complex: Potential for NMR as a Key tool in therapeutic antibody design and development. The Journal of Biological Chemistry.
- Holliger, P., Prospero, T., and Winter, G. (1993). “Diabodies”: small bivalent and bispecific antibody fragments. Proc. Natl. Acad. Sci. U.S.A. 90, 6444–6448.
- Iliades, P., Kortt, A.A., and Hudson, P.J. (1997). Triabodies: single chain Fv fragments without a linker form trivalent trimers. FEBS Lett. 409, 437–441.
- Ward, E.S., Güssow, D., Griffiths, A.D., Jones, P.T., and Winter, G. (1989). Binding activities of a repertoire of single immunoglobulin variable domains secreted from Escherichia coli. Nature 341, 544–546.
- Muyldermans, S., Atarhouch, T., Saldanha, J., Barbosa, J.A., and Hamers, R. (1994). Sequence and structure of VH domain from naturally occurring camel heavy chain immunoglobulins lacking light chains. Protein Eng. 7, 1129–1135.
- Greenberg, A.S., Avila, D., Hughes, M., Hughes, A., McKinney, E.C., and Flajnik, M.F. (1995). A new antigen receptor gene family that undergoes rearrangement and extensive somatic diversification in sharks. Nature 374, 168–173.
- Stanfield, R.L., Dooley, H., Flajnik, M.F., and Wilson, I.A. (2004). Crystal structure of a shark single-domain antibody V region in complex with lysozyme. Science 305, 1770–1773.
- Nuttall, S.D., Humberstone, K.S., Krishnan, U.V., Carmichael, J.A., Doughty, L., Hattarki, M., Coley, A.M., Casey, J.L., Anders, R.F., Foley, M., et al. (2004). Selection and affinity maturation of IgNAR variable domains targeting Plasmodium falciparum AMA1. Proteins 55, 187–197.
- Streltsov, V.A., Varghese, J.N., Carmichael, J.A., Irving, R.A., Hudson, P.J., and Nuttall, S.D. (2004). Structural evidence for evolution of shark Ig new antigen receptor variable domain antibodies from a cell-surface receptor. Proc. Natl. Acad. Sci. U.S.A. 101, 12444–12449.
- Wesolowski, J., Alzogaray, V., Reyelt, J., Unger, M., Juarez, K., Urrutia, M., Cauerhff, A., Danquah, W., Rissiek, B., Scheuplein, F., et al. (2009). Single domain antibodies: promising experimental and therapeutic tools in infection and immunity. Med. Microbiol. Immunol. 198, 157–174.
- Qiu, X.-Q., Wang, H., Cai, B., Wang, L.-L., and Yue, S.-T. (2007). Small antibody mimetics comprising two complementarity-determining regions and a framework region for tumor targeting. Nat. Biotechnol. 25, 921–929.
- Martens, T., Schmidt, N.-O., Eckerich, C., Fillbrandt, R., Merchant, M., Schwall, R., Westphal, M., and Lamszus, K. (2006). A Novel One-Armed Anti-c-Met Antibody Inhibits Glioblastoma Growth In vivo. Clin Cancer Res 12, 6144–6152.
- Rose, R.J., Labrijn, A.F., van den Bremer, E.T., Loverix, S., Lasters, I., van Berkel, P.H., van de Winkel, J.G., Schuurman, J., Parren, P.W., and Heck, A.J. (2011). Quantitative analysis of the interaction strength and dynamics of human IgG4 half molecules by native mass spectrometry. Structure 19, 1274–1282.
- Wilkinson, I.C., Fowler, S.B., Machiesky, L., Miller, K., Hayes, D.B., Adib, M., Her, C., Borrok, M.J., Tsui, P., Burrell, M., et al. (2013). Monovalent IgG4 molecules: Immunoglobulin Fc mutations that result in a monomeric structure. mAbs 5, 406–417.
- Ishino, T., Wang, M., Mosyak, L., Tam, A., Duan, W., Svenson, K., Joyce, A., O’Hara, D.M., Lin, L., Somers, W.S., et al. (2013). Engineering a Monomeric Fc Domain Modality by N-Glycosylation for the Half-life Extension of Biotherapeutics. J. Biol. Chem. 288, 16529–16537.
- Ying, T.L., Chen, W.Z., Gong, R., Feng, Y., and Dimitrov, D.S. (2012). Soluble Monomeric IgG1 Fc. J Biol Chem 287, 19399–19408.
- Hopp, J., Hornig, N., Zettlitz, K.A., Schwarz, A., Fuß, N., Müller, D., and Kontermann, R.E. (2010). The effects of affinity and valency of an albumin-binding domain (ABD) on the half-life of a single-chain diabody-ABD fusion protein. Protein Engineering, Design and Selection 23, 827–834.
- Cuesta, A.M., Sainz-Pastor, N., Bonet, J., Oliva, B., and Alvarez-Vallina, L. (2010). Multivalent antibodies: when design surpasses evolution. Trends Biotechnol. 28, 355–362.
- Nelson, A.L. (2010). Antibody fragments: hope and hype. MAbs 2, 77–83.
 United Kingdom (UK)
United Kingdom (UK)